2016 CSD Data and Software Updates
Here we describe the CSD data and software updates provided in 2016.
The Cambridge Structural Database Data Update – May 2016
We are delighted to announce the release of the May 2016 update of the
Cambridge Structural Database (CSD). With 14,996 new entries from 6,057 new
publications, this update brings the total size of the CSD to over 833,000
entries.
New data highlights include the structure of the protein tyrosine kinase
inhibitor regorafenib (refcode: QABCEE, DOI: 10.5517/ccdc.csd.cc1430hr),
where conformational differences in the single component structure of this anticancer
compound are compared to that of the monohydrate form. The update also contains a
fascinating series of crystal structures of the metal organic framework FJI-H11
(including refcode: UMUBAH, DOI: 10.5517/ccdc.csd.cc1kn2dp),
where the framework is seen to undergo extremely large structural expansion and
contraction in response to temperature and solvents.
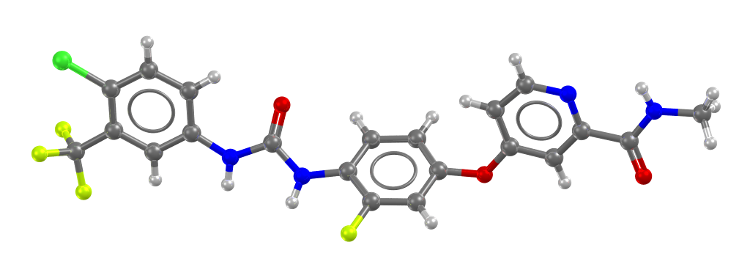
Potent anticancer compound regorafenib
(CSD Refcode: QABCEE, DOI 10.5517/ccdc.csd.cc1kn2dp)
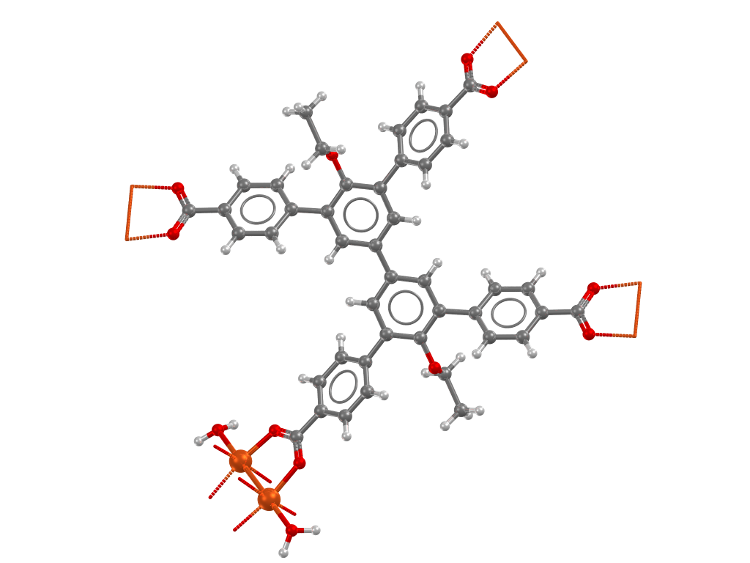
Structure of the metal organic framework
FJI-H11 which shows large structural transformations (refcode: UMUBAH,
DOI: 10.5517/ccdc.csd.cc1kn2dp)
The Cambridge Structural Database Data and Software Update – February
2016
We are delighted to announce the release of the
first 2016
update of the Cambridge Structural Database
(CSD). With 17,968 new entries from 7,231 new publications, this update brings the total
size of the CSD to over 818,000 entries.
New data highlights include a new polymorph of aripiprazole (refcode: MELFIT19, In addition to the data update we’ve continued our integration and Your CSD-System now includes easy access With direct links to our data deposition service through the CSD-System it is The ninth polymorph with a solved crystal A novel chromium complex with a unique On the software front, we’ve continued the integration of our The simplest way to update is to click on Help->Check for Updates in We hope you enjoy your new
DOI: 10.5517/cc1jqyz5), the most
polymorphic drug published to date, with a record-breaking ninth solved crystal
structure. The update also contains a fascinating chromium complex (refcode: XUZLUB,
DOI: 10.5517/cc1jyy6m
development of CSD-based software. Mercury now acts as a hub for the huge range of such
applications and services – but of course all can still be used in stand-alone
mode.
to CSD-Community,
the set of essential software and web services freely available to all scientists.
Crystallographers can now deposit structures directly from the Mercury
interface, teachers can access our carefully selected CSD Educational Collection of over
700 structures and of course anyone can access data from the CSD online and use our
community tools such as enCIFer.
now even easier for you to publish and share your structures directly through the CSD.
In 2015 we enhanced our popular Private Communications system to
deliver CSD
Communications. Last year over 4,000 structures were published in this way and
in 2016 a CSD
Communication looks set to be the most
popular method to publish crystal structures. Such structures are given a citable DOI,
links from third party repositories and indexes such as the Thomson Reuters Data
Citation Index and of course get a thorough review by our expert scientific editorial
team before entering the CSD. We believe that scientists should be able to share their
data without financial barriers – A CSD
Communication is therefore both free to
create and free to view.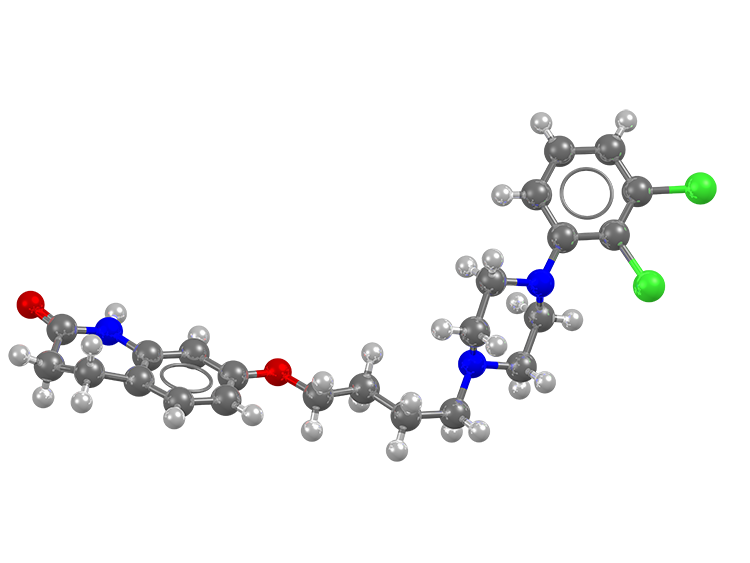
structure of aripiprazole (refcode: MELFIT19,
DOI: 10.5517/cc1jqyz5)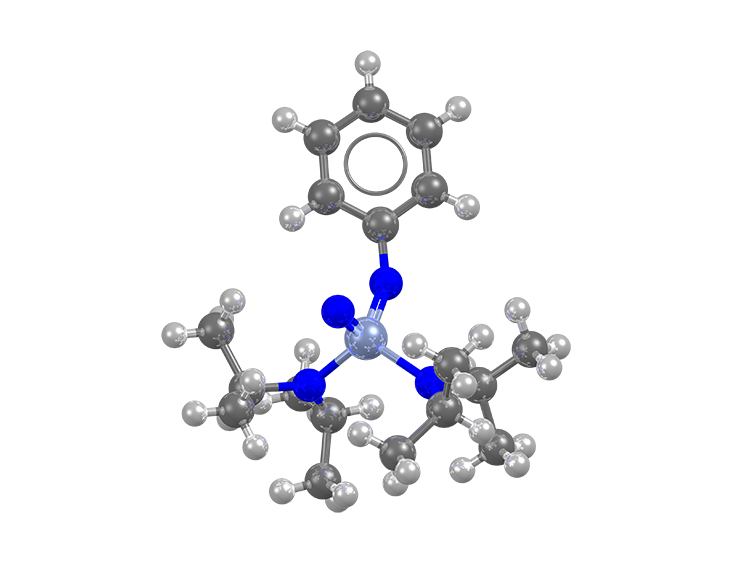
bonding pattern (refcode: XUZLUB, DOI: 10.5517/cc1jyy6m)
applications into CSD-Enterprise. All our academic community and our Research Partners
will see CSD-Community,
CSD-System, CSD-Materials,
and CSD-Discovery functionality
sitting alongside each other, complemented by a menu for your own Python scripts,
allowing you to access and link CSD functionality in any way you imagine. Of course,
we’ve started you off with some scripts developed at the CCDC and
the CSD Python API
Forum provides a place to both pick up new
ones and share your own.
Mercury or Hermes. CSD-Enterprise is strong stuff; if you use Windows 7 or above
right-click on Mercury and select the ‘run as administrator’ option for this update
process. Restart Mercury and you’ll see the improvements we’ve
made.
structures and software.