SuperStar
Visually Probe Protein-Ligand Structures and Understand Their Binding Sites
Providing knowledge-based pharmacophore generation and prediction of intermolecular interactions, SuperStar is used by medicinal chemists to visualize and understand molecular design by protein interaction mapping.
SuperStar uses crystallographic information about non-bonded interactions to generate interaction maps within protein binding sites or around small molecules, i.e. it predicts ‘hot-spots’ where a chosen interaction type is exceptionally favourable.
The interaction maps are generated by estimating the probability of an interaction between, for example, the protein and a probe (a small functional group such as methyl or carbonyl) based on how often the interaction has been observed in crystal structures.
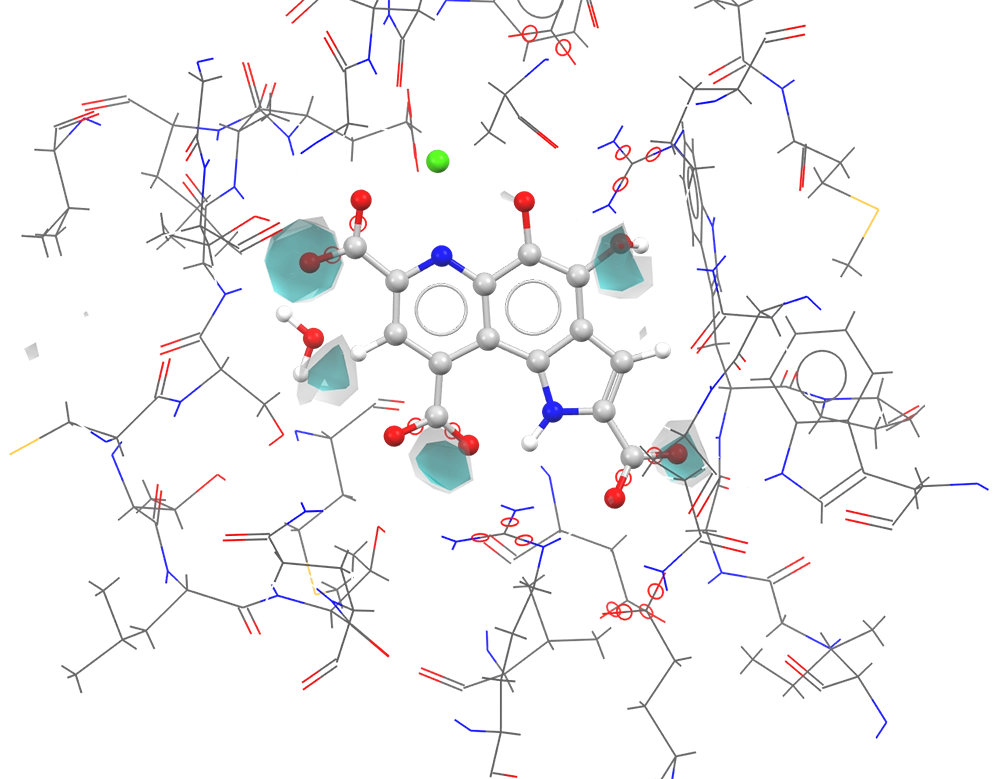
SuperStar contour map of 4AAH (Methanol Dehydrogenase) binding site using the C=O probe (CSD data). The ligand acceptor groups correlate well with the areas of highest predicted propensity.
Benefits
Simple interface
Fast, simple wizard to run the analysis.
Expertly-presented results
Interpret the outputs and guide your molecule design. Likelihoods of interactions rather than comparative energy - this is probably most important for weaker probes.
Real data
Based on real experimental data from the CSD. Complementary to calculated methods like DFT, SAPT. Competition information is implicit in the underlying data.
Accessible via Python API and Hermes desktop GUI
Custom control or run on many proteins in a workflow, and use outputs for machine learning.
Time efficient
Faster than using energy-based calculations.
Full Interaction Maps (FIMs)
Map interaction preferences around complete molecules in a crystal structure. Visualise observed atom-atom contacts with respect to likely geometries in 3D space. Identify interaction hotspots around chemical groups.
Included in
Fields
Use Cases
Learn more
See our complete and up-to-date list of technical FAQs in our knowledgebase at the link above.
FAQs
SuperStar is knowledge-based and not based upon any forcefield. It draws on knowledge from experimentally determined crystal structures in the Cambridge Structural Database.
Superstar analyses proteins, giving a map of potential binding sites. FIMs is used to analyse small molecules, for example, to understand if H-bonds are in optimal locations.
SuperStar retrieves its data from IsoStar, CCDC’s library of intermolecular interactions. IsoStar contains information about non-bonded interactions from both the CSD and the PDB. The interaction maps that SuperStar generates are therefore fully knowledge-based and all results can be traced back to experimentally observed geometries of interaction.