Ending the Pesticide Drought using Research Techniques well established in the Pharmaceutical Industry
Techniques used in modern drug discovery are now being adopted by pesticide scientists to turn the tide in the pesticide resistance battle, with new modes of action being investigated and developed. In production, new software to predict and overcome manufacturing bottlenecks is equally beneficial to both the pharmaceutical and agrochemicals industries.
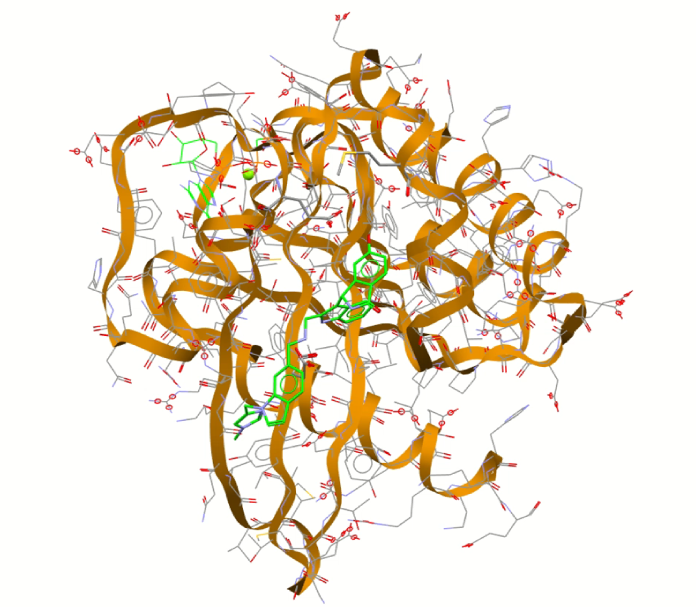
Protein-ligand docking—a common drug discovery technique now being used by the agrochemical industry to discover new pesticide candidates
The Battleground
Highly effective pesticides emerged on to the market in the 1970s and 1980s that were so successful that it was deemed not a good use of resources by many in the industry to carry on researching alternatives, or even understanding modes of action.
Take, for example, glyphosate-based herbicides that are used to suppress weeds. When glyphosate resistant crops were introduced, farmers could apply glyphosate herbicides widely with high effectiveness. The weeds were killed or suppressed; the crop plants thrived. Owing to this success, little attention and investment were given to pesticide research to find alternatives that at the time were deemed unnecessary.
The Drought and Counterattack
Since 1984, only one herbicide with a new mode of action has come onto the market, compared to once every 2 years on average during the previous 3 decades.[1]
The weeds (simply a plant in the wrong place at the wrong time) took full advantage of this period of complacency. During this time, instances of herbicide resistance increased 600%.[1] Today some farmers are now having to change the crops they grow owing to a lack of effective weed-suppressing herbicides.
New Research
With a newfound urgency, attention and investment has returned to pesticide research, with scientists turning to techniques used in other industries, including the pharmaceutical industry with its established drug discovery process. New software to optimise manufacturing and processing is equally applicable to both industries.
What Are Pesticides and How do They Work?
Pesticides are toxic substances used for suppressing or killing animals, plants or fungi that are detrimental or harmful to cultivated plants or animals.
Often hazardous to the health of domestic animals, humans and wildlife, pesticides interfere with the normal metabolic processes in the pest (target) organism. They are classified by the organism that they are targeted at:
- Herbicides – inhibits weeds and invasive species (for example glyphosates for the treatment and control of Japanese Knotweed; sulfonylurea for the treatment and control of grasses in crops).
- Insecticides – kills insects that infest and/or carry disease (for example ketoenols for the control of aphids).
- Fungicides – kills or inhibits the growth of fungi that cause crop damage or endanger the health of animals (for example azoles for the prevention of rust and mildew in crops).
- Fumigants – gaseous pesticides to suffocate or poison insects and other organisms that damage stored foods, houses, and cloths (for example sulfuryl fluoride used in termite ‘tenting’ in houses with an infestation).
Pesticides have several routes to penetration:
- Ingestion – for example pesticide is sprayed onto foliage that is then eaten by caterpillars and grasshoppers.
- Inhalation – for example fumigants for killing pests in stored products.
- Contact – penetration of body covering, for example aphids.
During the pesticide ‘drought’, pests have evolved to avoid these modes of action.
Drug Discovery Techniques to the Rescue
Pesticide scientists are now turning to techniques used in the pharmaceutical industry – protein-ligand docking used in drug discovery – to look at new ways to protect crops.
In drug discovery, new drugs are developed using techniques to dock and bind ligands to the protein surfaces of disease-causing organisms in silico (on a computer), to see which compounds inhibit the protein, killing the target organism, e.g. a virus. Millions of potential compounds are screened against macromolecular targets using software to discover new drug candidates.
Software tools in the CSD-Discovery suite, drawing on the knowledge obtained from 1.1M+ crystal structures held within the Cambridge Structural Database (CSD), are used by pharmaceutical scientists to discover new drug molecules by:
- Validating geometries including bond lengths and angles
- Identifying alternative conformations
- Analysing, predicting and understanding protein and ligand interactions
- Identifying common interaction patterns with data mining
Discovery scientists can perform protein-ligand docking experiments and virtual screening with GOLD, the validated, configurable protein-ligand docking software for expert drug discovery, including virtual screening through to lead optimization.
Included in the CSD is a Pesticide Subset of nearly 1000 entries linked directly to the Pesticide Property Database (PPDB) produced by the Agriculture & Environment Research Unit (AERU) at the University of Hertfordshire. The subset allows users to browse and restrict searches to the pesticide entries.
The protein-ligand docking mode of action can be applied to new herbicides that could work in a similar way – binding to one of the many enzymes critical to the plant’s survival.[2] Research is now focussing on identifying enzymes which are critical to the unwanted plant’s survival that have large deep pockets suitable for binding, and then finding herbicide candidates that will bind to these pockets and kill the plant.
These new herbicides must also be environmentally safe (taking learnings from the DDT environmental disaster[3]), harmless to humans, and cheap.
Predicting and Overcoming Manufacturing and Processing Obstacles
In the pharmaceutical industry, processing problems of crystalline products include issues in flow, wettability, tabletability, and stickability. These problems can make materials hard to process, and are typically discovered by resource-intensive manufacturing trials. Extra processing steps, more frequent clean-downs, or alternative formulations are then used to overcome them.
These issues also apply to agrochemical manufacture and the techniques used by the pharmaceutical industry to predict and overcome them are equally transferable and applicable.
Drawing on data from the CSD, the new CSD-Particle suite from the CCDC can analyse the mechanical and chemical properties of crystalline particles, e.g. pharmaceuticals and agrochemicals. This data and insight can then be used to find the molecular cause of manufacturing issues to assess risks, anticipate bottlenecks, and identify problematic formulations before costly manufacturing trials, allowing researchers to eliminate many candidates and focus on those predicted to have minimal manufacturing issues.
“This knowledge will be pivotal to improving development timelines for pharmaceutical products in the future,” said Dr. Stephen Thomas (Modeling Scientist) and Dr. Amy Sarjeant (Associate Scientific Director), Bristol Myers Squibb, NJ, USA.
Next Steps
Unlock the knowledge from over 1.1M crystal structures in the Cambridge Structural Database (CSD).
Develop new modes of action including protein-ligand docking with CSD-Discovery.
Predict and overcome manufacturing and processing obstacles with CSD-Particle.
References
[1] https://cen.acs.org/environment/pesticides/crop-protection-herbicide-mode-action-glyphosate/100/i22.
[2] See GOLD in action: leveraging structure-based drug design methods to develop novel herbicides - The Ca mbridge Crystallographic Data Centre (CCDC).