Faster, Easier Custom Pharmacophore Queries
What’s New in CSD‑CrossMiner
Pharmacophore searching in CSD-CrossMiner has been improved by simplifying the creation of chemically precise features and making it easier to save, reuse, and refine search results as ideas develop.
Saving and Reloading Sessions for Analysis and Sharing
CSD-CrossMiner now supports saving and loading sessions using File > Save Session… and File > Load Session…
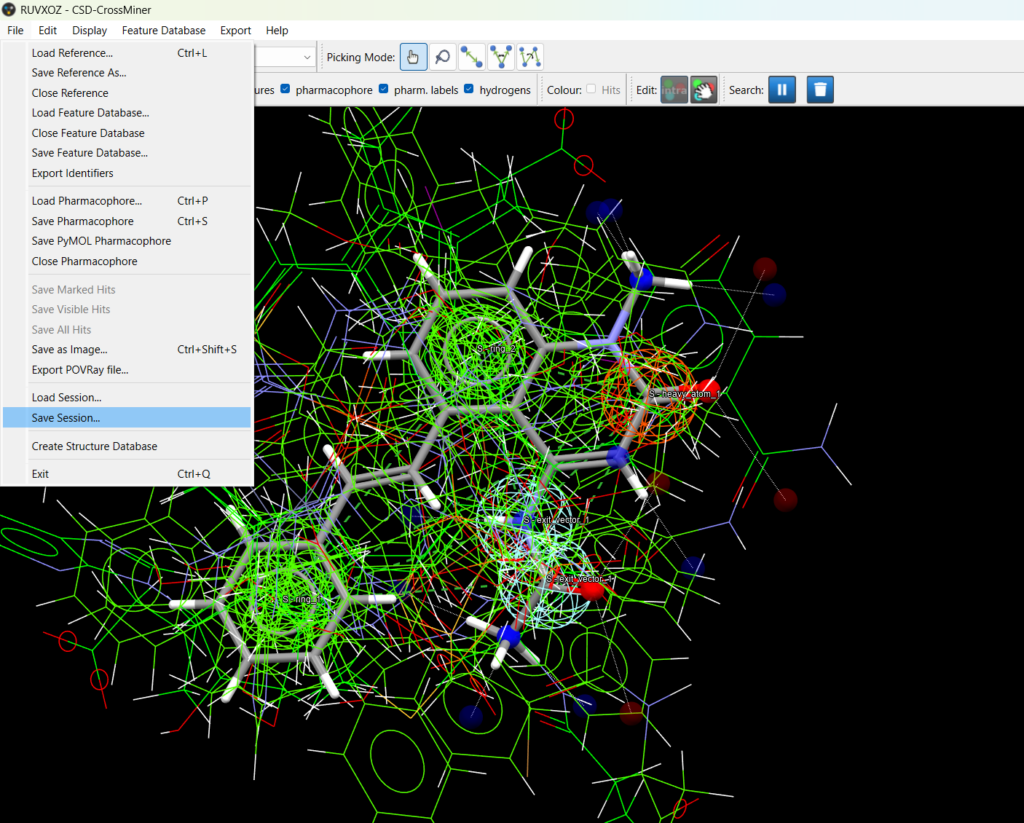
A saved session, stored in a binary .cmx format, captures:
- the pharmacophore query
- the reference structure
- all search results obtained up to the point at which the session was saved.
Sessions can be saved even while a search is still running. When a session is reloaded, the results present at the time of saving are restored exactly as they were and can be inspected and analysed further.
This functionality enables users to compare different query variants by saving sessions at key decision points and step-by-step updating queries without losing the results of previous searches. Before making a small change to a query, a session file can be saved to create a checkpoint, preserving a meaningful set of results for each refinement step.
Session files also simplify sharing a complete pharmacophore setup and its associated results with collaborators for discussion or review. The only requirement is that CSD-CrossMiner has access to the same feature and structure databases that were used when the session was saved. If these databases are not found at their original locations, the user is prompted to locate them before loading proceeds.
It is important to clarify that searches cannot currently be resumed from a saved session. Clicking the Start pharmacophore search button after loading a session will always restart the search from the beginning and replace the stored results.
Sketch‑Based Definition of SMARTS
SMARTS patterns are used throughout CSD-CrossMiner in several contexts to define chemical substructures. Manually writing SMARTS can become hard to manage, particularly as definitions grow more complex. To simplify substructure definition, an interactive structure sketcher has been introduced into CSD-CrossMiner and can be used wherever SMARTS definitions are required.
The sketcher allows a substructure to be drawn graphically and automatically converted into a SMARTS pattern, which is applied immediately. Manual SMARTS entry remains available, but the sketcher makes it easier to build and modify detailed definitions without working directly with SMARTS syntax.
The sketcher interface is the same as that used in other software in the CCDC portfolio, such as ConQuest and Mercury. Its extensive functionality makes it easier to design substructures that would be tedious or error‑prone to specify manually.
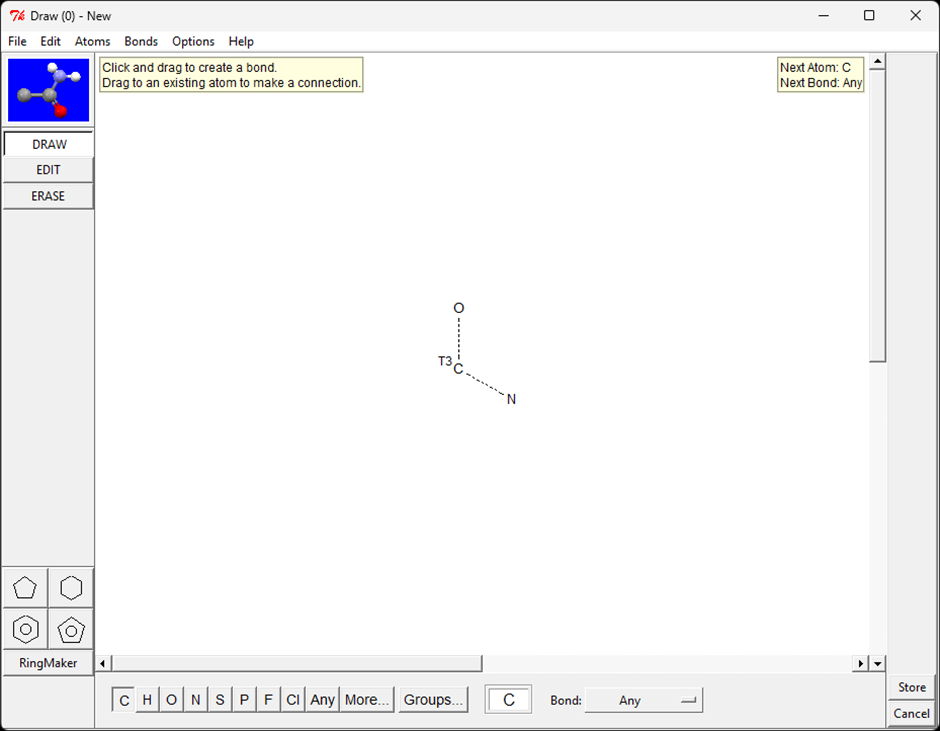
The sketcher is available in three contexts:
- Defining custom pharmacophore features
When defining new pharmacophore features in the Feature Editor, a new Sketch button is available. Clicking this button opens the sketcher and allows a substructure to be drawn. Clicking Store in the sketcher window converts the drawing into SMARTS, automatically assigns indices to all atoms, and adds it to the list of substructure definitions in the Feature Editor. If not all atoms in the substructure should be selected via the indices, the indices field can still be modified manually. Once the feature definition is finalised, selecting File > Add Feature to Current Feature Database adds the custom feature to the list of available pharmacophore features.
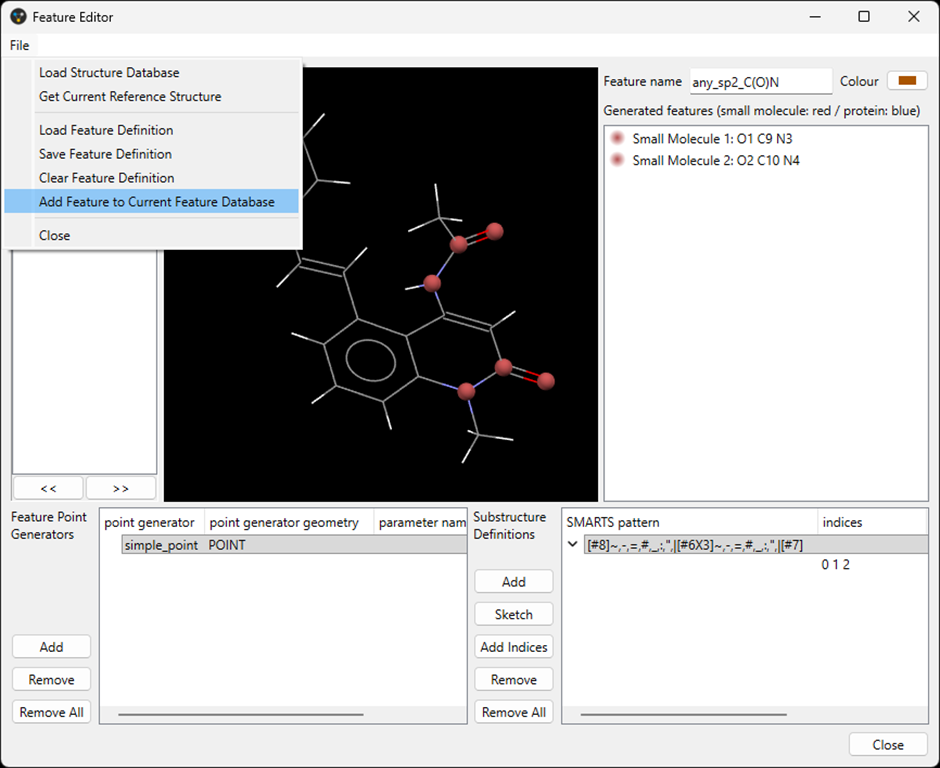
- Defining bespoke excluded volumes
Excluded volumes define regions of space where certain atoms cause a hit to be rejected. While excluded volumes default to excluding any atom, SMARTS patterns can be used to make exclusions chemically selective. The Excluding atoms (SMARTS) field now supports sketch‑based input (by right‑clicking and selecting Sketch), making it easier to target specific atom types or functional groups. - Creating substructure filters
Substructure filters constrain pharmacophore searches by requiring the presence or absence of a specific chemical motif. The SMARTS field of a substructure filter now also supports sketch‑based input by right‑clicking on the field and selecting Sketch.
Additional Improvements
There have also been several general usability improvements to CSD-CrossMiner.
Intramolecular constraints are now enabled by default, and any new Small Molecule pharmacophore point is automatically added to the set of constrained points. This reflects the most common use case and helps ensure that typical small‑molecule queries behave as expected out of the box. Full manual control over constraints remains available once all pharmacophore points have been added.
In addition, drag‑and‑drop onto the application window is now supported for pharmacophore queries (.cm), reference structures (common molecular file formats), feature databases (.feat), and saved sessions (.cmx).
Next Steps
- Learn more about CSD-CrossMiner to build pharmacophore queries and simultaneously mine the CSD, PDB, and your proprietary structural data to quickly identify off-target effects, alternative scaffolds, similar binding sites, interaction motifs, bioisosteres, and more.
- CSD-CrossMiner is part of CSD-Discovery for computer-aided drug discovery.
- Want to see these updates live? Join our webinar on 21 May and discover what’s new in CSD‑CrossMiner. Register here.
- To discuss further and/or request a demo with one of our scientists, please contact us via this form or email us at hello@ccdc.cam.ac.uk.