Using the CSD Python API with Excel
Love it or hate it, Excel is extensively used in industrial settings. Often folk like to use Excel to analyse the results of their work generated from some script or workflow. The usual workflow is to dump out a CSV file and load this into Excel, but sometimes this is somewhat inefficient. (How many times have you dumped a CSV file, loaded it, re-filtered it etc. etc. only then to discover that maybe the output needs one extra field you should have added, so you end up repeating the analysis run with a small script change and then going through all the steps on loading the Excel file again. If you end up doing this a lot, it gets inefficient!).
Fortunately, Python3 has modules that allow direct interfacing. There’s an open-source package called xlwings and a commercial package called PyXLL. In this blog we will look at using xlwings, but we note that one could achieve the same results using PyXLL instead. The two technologies differ in xlwings and PyXLL. PyXLL is a full add-in to Excel and consequently doesn’t rely on the user having to have macros enabled in their spreadsheets. xlwings is nice for personal spread sheets using Macros whereas PyXLL is probably better for rolling out a standard set of functions, which might be useful for deployment to (for example) a team of Excel savvy Medicinal Chemists.
In this short blog, we’ll explore how we can use User Defined Functions to add in macros into Excel that call the CSD Python API.
Prerequisites
Firstly, you’ll need to get xlwings and the python3 environment installed. This was how I did this on my PC. Note – because the CSD Python API with Python 3 distribution is 64 bit, I used the recommended 64 bit version of Excel. It was however also possible to get the examples to work with 32 bit Excel too.
- Install the appropriate version of Anaconda (For me this was 64 bit Anaconda3)
- Start an Anaconda powershell
- create a python 3.7 environment (note here I used the command
conda create -n test_xlwings python=3.7) - type
conda activate test_xlwings - in the power shell, type xlwings addin install
- Download the apprpriate csd-python-api zip file and unpack it (e.g. to
E:csdpythonapiccdc_conda_channel) - Install the csd-python-api into the your conda environment (
conda install -c <Where-ever the CCDC conda channel is - e.g. E:csdpythonapiccdc_conda_channel> csd-python-api) - change to a directory you want to work in in the powershell
- type in the powershell
xlwings quickstart my_project - This will create a folder called my_project – go into it. In the folder there will be two files: one called
my_project.xlsmand another calledmy_project.py
The XLSM file is a macro-enabled excel file. The file my_project.py is a stub file containing a basic example of using xlwings.
So, you can now create nice functions that allow you to embed python code as functions into Excel. The my_project.py file contains a default example; it looks like this:
The first function in this file is an example of a plain callable function that can be called from VBA. We won’t be using this in this demonstration, but if you are interested you can go and read the xlwings quickstart documentation here. For our purposes, let’s delete this function.
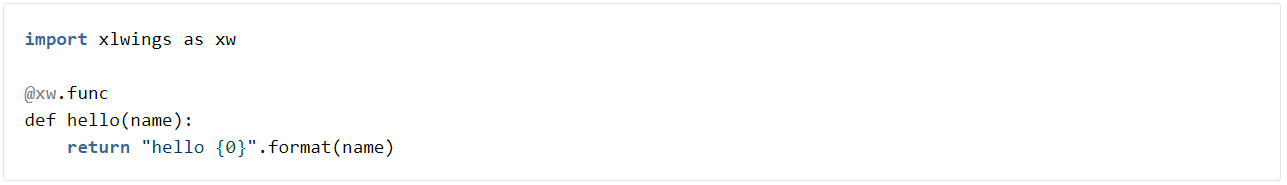

The ‘hello’ function is an example of a User Defined Function and it can be called from within Excel as a macro. That’s rather neat, and it lends itself to all sorts of extensions that you may find useful. We won’t repeat the xlwings documentation in detail here, but you can learn more about VBA UDFs here. Note that you can achieve the same things using PyXLL. For information on that see here.
To use the above function is easy. You open up my_project.xlsm in Excel. You should have an xlwings tab:

Clicking the “Import Functions” button will include the function ‘hello’ and so you can use it in a spreadsheet just like you’s use a built in Excel formula:
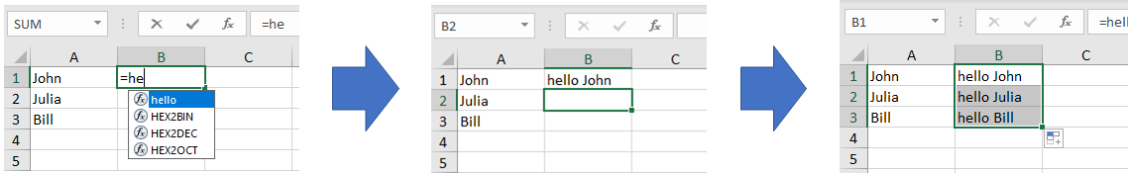
So far, we haven’t actually done any chemistry, but lets have a go. In this blog we are going to try two different things. Firstly, we are going to access the CSD and get some properties of entries displayed directly in a spread sheet. We will then write some extensions to allow us to filter the entries based on substructures. In the second example we will look at creating a dynamic functions to perform searches across the whole CSD.
Example 1. Accessing Properties in the CSD
A really simple thing we might want to do as a user is to have a list of properties for an entry in the CSD. This is pretty simple using a UDF function. All we need to do is open the CSD as an entry editor and then create a UDF function that accesses the property. For example the following function will retrieve the chemical name for entries in the CSD.
To try this out, firstly go to the my_project folder and edit my_project.py.
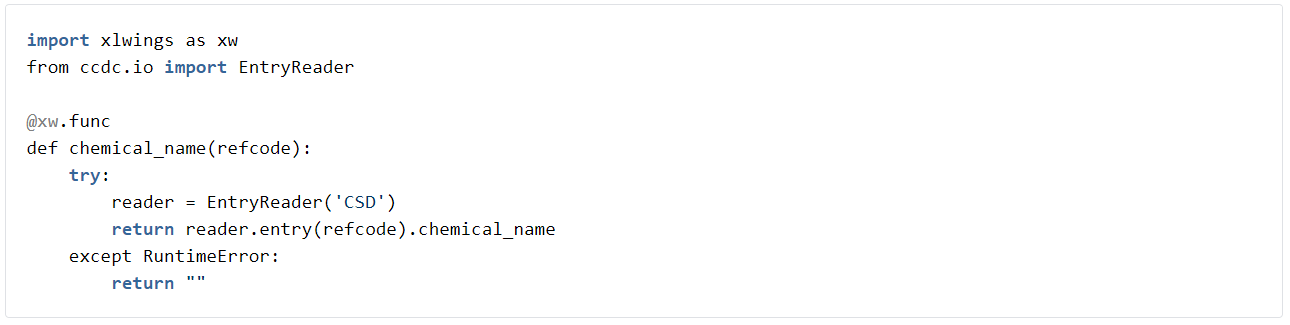
Now try opening my_project.xlsm in Excel. Create a table in Sheet1 like the sheet shown below
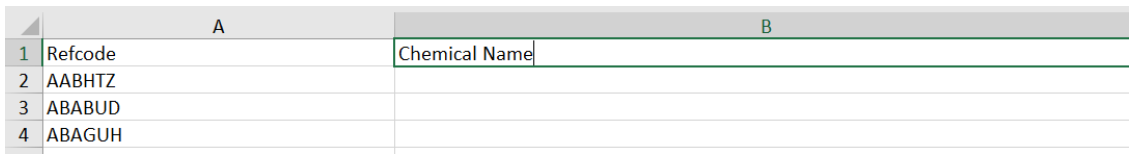
You can now type into cell B2 and select the =chemical_name() function … this is the UDF function we just created in the python code above
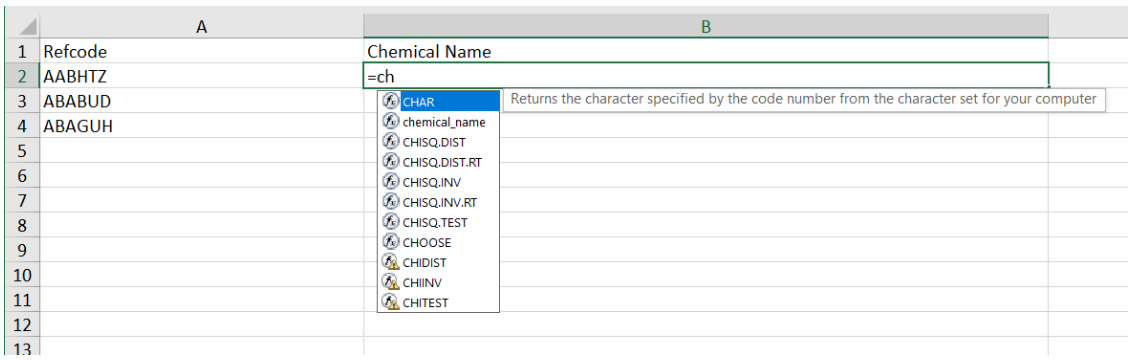

Selecting chemical_name and adding in cell A2 as below
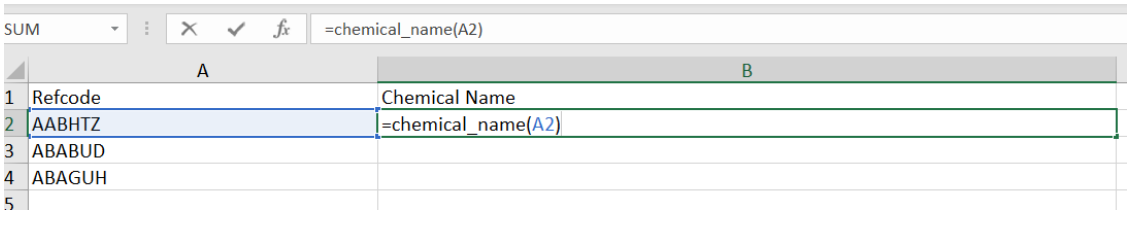

then hitting return then gives the following: as you can see the chemical name for the entry in AABHTZ has been returned.
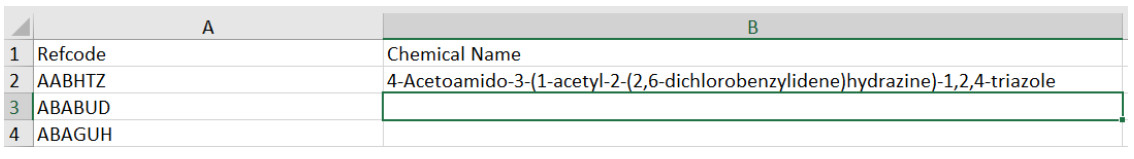

Of course, you can then, as standard in Excel drag the formula and populate all the cells.
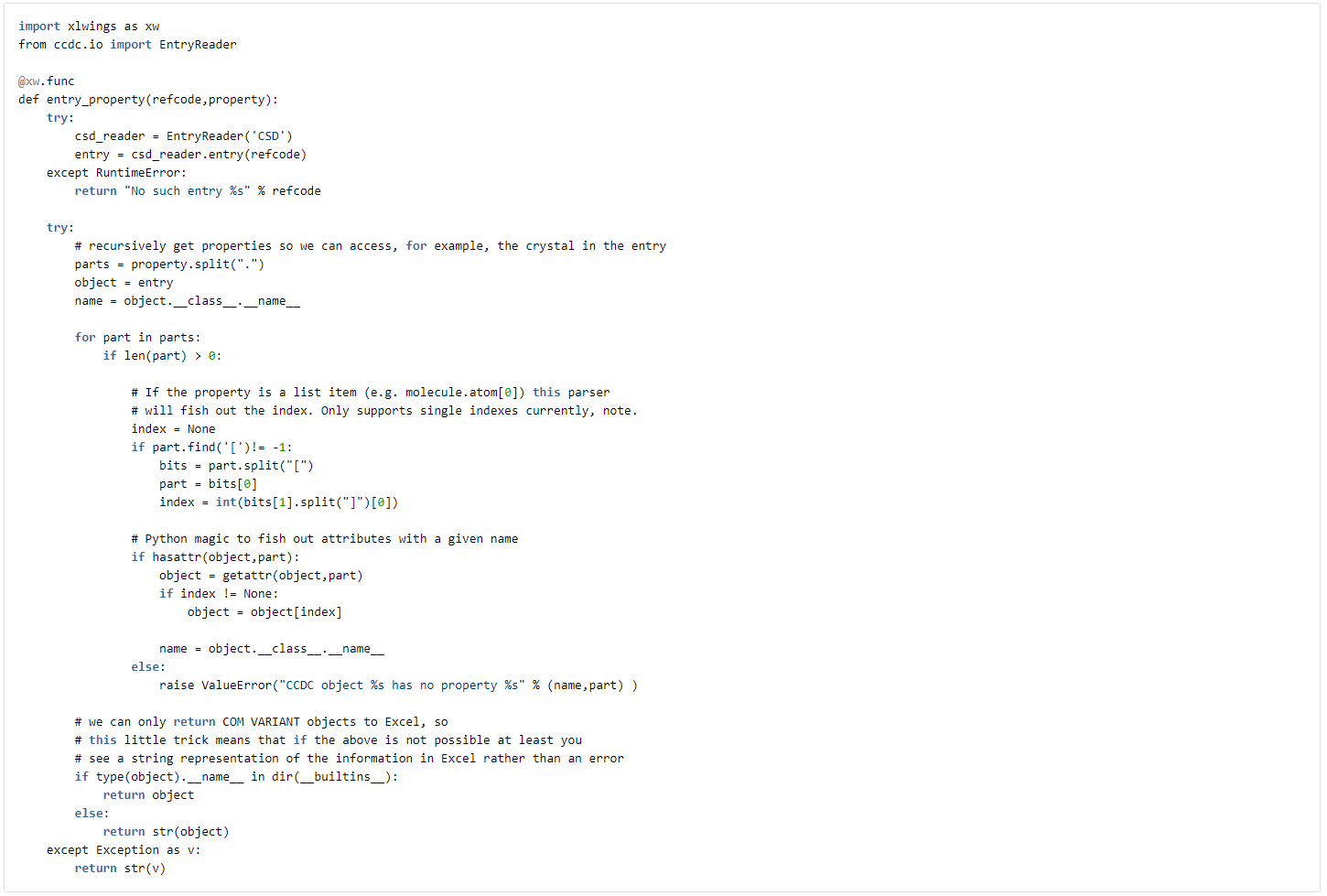
A far more elaborate example allows us to access pretty much any property from a CSD entry by using some python magic

In the above example, we can, for example, type =entry_property(“AABHTZ”,”molecule.smiles”) and retrieve the SMILES code for the complete CSD Entry. In fact, if you know the python API syntax you can access all the attributes on an entry using the approach above.We might want to consider filtering on substructures. A simple example below allows us to create a filter function that will search for an individual substructure in a given structure

You can then use this simple formula to count how many occurrences there are of a specific substructure in a range of CSD entries: for example, you might type =substructure_count(“AABHTZ”,”c1ccccc1″) into a cell to count the number of phenyl rings in AABHTZ.
The above examples show how easy it is to use a combination of xlwings and the CSD Python API to extend excel for extracting molecular properties. In example 2 we will see how we can also do full searching using the API and array functions.
Example 2. Using an Array Formula to perform searches across the whole CSD
It is possible to do full searches of the CSD using an array formula. In this example, the code will generate a list of refcodes in the CSD that contain a specific line of text and insert a column of refcode into the spreadsheet. A similar approach is also available for SMARTS driven substructures and Similarity searching in the CSD.
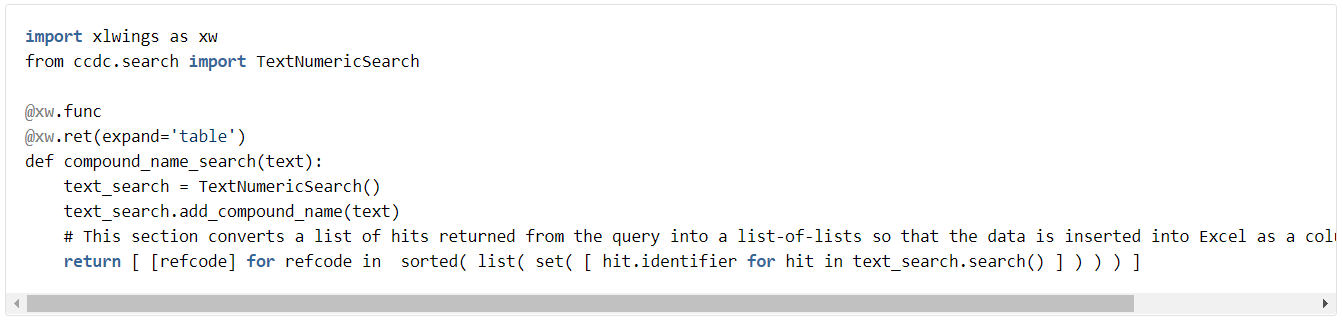
xlwings has a useful code ‘decorator’ to control the return type; @xw.ret(expand=’table’). This allows the user to generate array functions that return 2D arrays of data. In the first example, lets do a text search that searches compound names. The function is added below. Note that we have to do some data manipulation at the end of the function to create a list of unique refcodes to return.
In excel, you can then find entries in the CSD that have bicyclononane in the title:
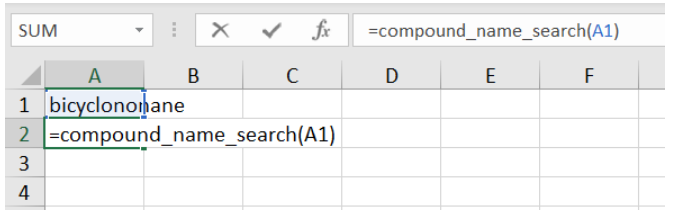
hitting return will then populate the Excel table with the structures that have bicyclononane in their chemical name

You can also do substructure searches in the same way. Below is an example of a simple SMARTS driven substructure search function that will search for a given SMARTS code, again returning a column of data:


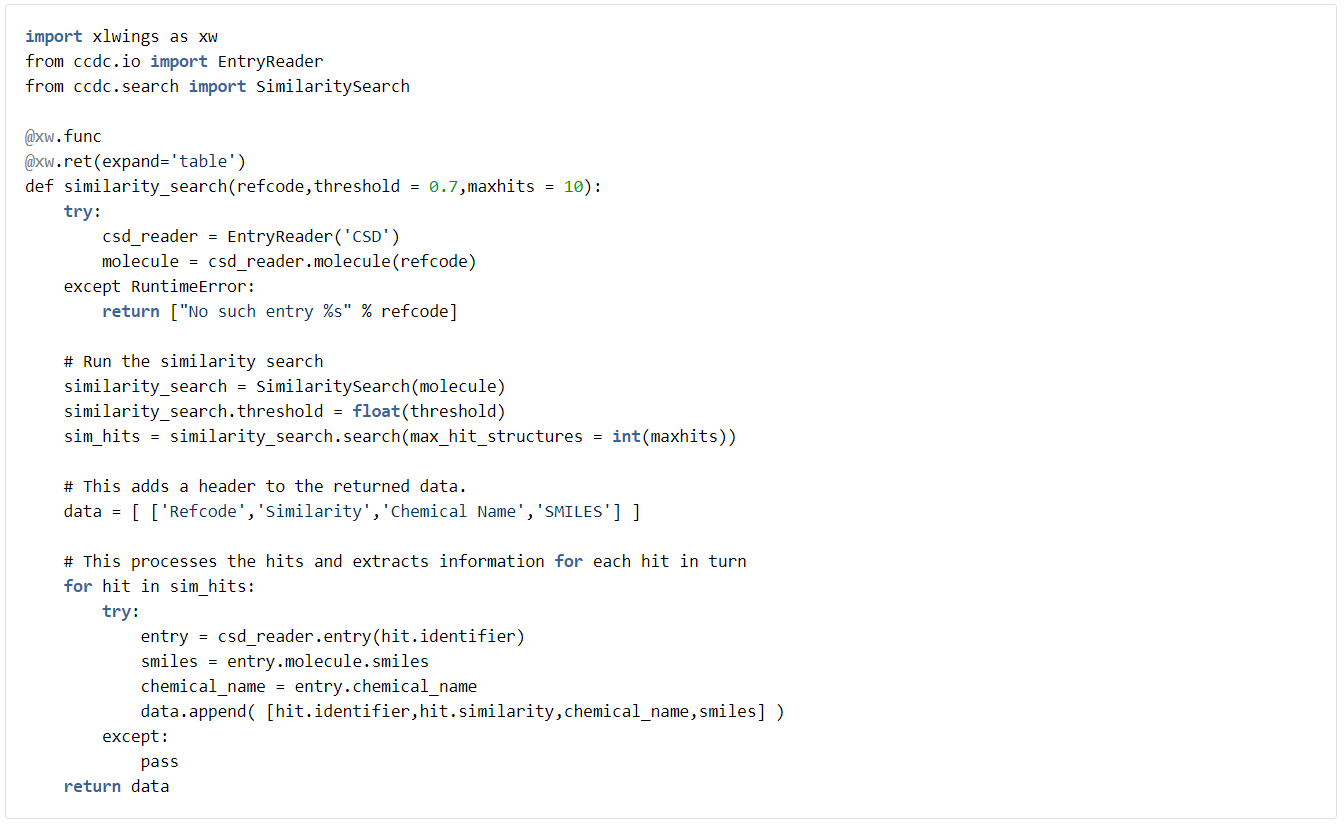
Finally, an example of a similarity search. In this case we will return multiple data items for each hit. The search looks for the 10 most similar structures in the CSD to a given refcode.
We will return the similar compounds’ refcodes, the similarity score, and their chemical names and SMILES codes:

If you’d like to see these examples in action, you can download the code here.