Pharmacophore based search – generating new ideas in drug discovery
Generating new ideas or looking at protein-ligand interactions in a novel way is key to innovative drug discovery and design. Pharmacophore models are an excellent way to do this.
This blog explores pharmacophores, their role in drug discovery, and how to perform a pharmacophore based search.
What is a pharmacophore?
A pharmacophore is a spatial collection of steric and electronic features which are necessary to a molecule’s interactions.
Put simply; a pharmacophore maps out why ligand X binds to protein Y in terms of fundamental chemical characteristics.
By viewing the protein-ligand binding in these terms, it becomes easier to find alternative ligands that would achieve the same end result.
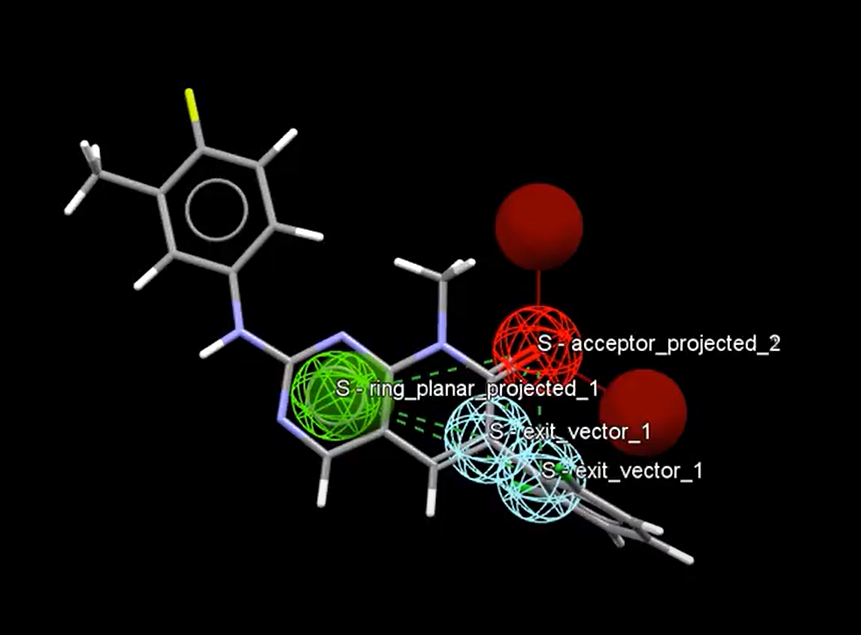
The pharmacophore of a molecule maps out the key electronic and steric features that are essential to its interactions. Here spheres mark out the key features on a molecule in CSD-CrossMiner.
How can pharmacophore search support drug discovery?
Viewing ligand-protein binding as its fundamental features in a pharmacophore, we can;
- Remove human bias or preconceptions
- Automate searches for alternative molecules
- Spot patterns of interactions which may help or hinder our work
- Identify possible cross-reactivity (other proteins that may bind our ligand)
- Search for alternatives to a natural ligand, where no protein structure is available (e.g. when a target protein is disordered)
How to perform a pharmacophore based search?
It’s possible to create and edit a pharmacophore then search against databases of structures using CSD-Crossminer.
By default you can search the Protein Data Bank and the Cambridge Structural Database – but you can also search in-house databases where required.
Learn how to set up and run a pharmacophore based search in this 5 minute video.