CSD Educators: Christine Zardecki from the Protein Data Bank (PDB)
For this CSD Educators blog, our guest is Christine Zardecki, from the RCSB PDB. This year, the Protein Data Bank (PDB) is celebrating their 50th anniversary and we are delighted to share with you a bit about their journey in education and the building of the PDB-101 educational resources.
Evolution of PDB-101: Molecular Explorations Through Biology and Medicine
Following the lead of Dr. Olga Kennard and the Cambridge Crystallographic Data Centre (CCDC), the structural biology community established the single worldwide archive for macromolecular structure data – the Protein Data Bank (PDB), in 1971 at Brookhaven National Laboratory in collaboration with the CCDC. From its inception, the PDB has embraced a culture of open access, ensuring its widespread use by the research community.
Today, the PDB archive is managed by the Worldwide Protein Data Bank (wwPDB), a consortium of organizations that host deposition, annotation, and distribution centers for PDB data from crystallographic (MX), electron microscopy (3DEM), and nuclear magnetic resonance (NMR) structural studies. wwPDB data centers include RCSB PDB (US), Protein Data Bank in Europe, and Protein Data Bank Japan BioMagResBank (US) archives related NMR spectra and related quantitative data.
Visit RCSB PDB and wwPDB for information about celebrations and more.
This year, RCSB PDB and wwPDB are celebrating the golden anniversary of the PDB with symposia, events, and related resources (ndr: see links above).
2021 marks another milestone important to RCSB PDB: the 10th anniversary of PDB-101 , RCSB PDB’s online portal for teachers, students, and the general public to promote exploration in the world of proteins and nucleic acids.
Building a Collection of Educational Resources
RCSB PDB became responsible for management of the PDB archive in 1998. Since January 2000, a Molecule of the Month (MotM) column by David S. Goodsell (RCSB PDB-Rutgers and The Scripps Research Institute) has been published regularly. This popular series tells stories about PDB structures, focusing on their functions and diverse roles within living cells, and highlights growing connections between biology, medicine, and nanotechnology. Monthly content has inspired readers around the world, and is a regular read for students and researchers alike (interested readers can subscribe for publication alerts here). Molecule of the Month articles are also known for their vivid illustrations; most are generated by molecular graphics programs, but several feature water color paintings that provide the visual context for molecules inside the cell.
All Molecule of the Month content, along with any materials developed by the RCSB PDB, are available free for use and reuse.
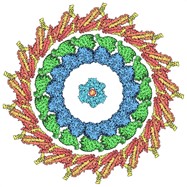 A kinetochore ring (yellow and red) surrounding a kinesin-microtubule complex (green and blue), and PCNA (turquoise) surrounding DNA (red, at center) from Twenty Years of Molecules. RCSB PDB plans to highlight this anniversary throughout 2020 were replaced by a focus on Coronavirus.
A kinetochore ring (yellow and red) surrounding a kinesin-microtubule complex (green and blue), and PCNA (turquoise) surrounding DNA (red, at center) from Twenty Years of Molecules. RCSB PDB plans to highlight this anniversary throughout 2020 were replaced by a focus on Coronavirus.
This series, which links to corresponding examples of PDB structures, was the main focus of RCSB PDB online educational development for many years. Progressively, more online materials were added to the collection of “educational resources” section of RCSB.org. These include the first edition of the Molecular Machinery poster (2002) that provides a tour of PDB structures and a series of Education Corner articles published in the RCSB PDB quarterly newsletter. This collection offers an account of how members of the community use the PDB to educate students, and includes CCDC-authored features about Using 815,527 Crystal Structures to Teach Chemistry: The Cambridge Structural Database (2016) and Symposium on the Applications of Small-molecule Crystal Structure Information in Chemical Education (2009).
The trove of educational materials continued to grow with resources on How Do Drugs Work? and Looking at PDB Structures (now Guide to Understanding PDB Data). With more than 100 Molecule of the Month features available, RCSB PDB had established an archive of educational content. However, features were not easily accessible unless visitors were familiar with PDB structures or knew the specific molecule of interest. Thinking like a data archive or database, RCSB PDB assigned categories to Molecule of the Month articles (Protein Synthesis, Enzymes, Health and Disease, Biological Energy, Infrastructure and Communication, and Biotechnology and Nanotechnology) so users could approach the archive with a question (how are genes replicated?) rather than an answer (DNA polymerase).
Launch of PDB-101
This top-down approach to packaging educational content led to the first release of the educational online portal PDB-101 in 2011. The name “101” denotes an entry-level course and is meant to signal the intended users: high school and undergraduate students. Our goal was to provide an “E-Z PDB” (one of the considered names). The main focus was the Structural View of Biology browser which centered individual Molecule of the Month articles into a greater, categorized molecular context. Branding all RCSB PDB education content under this highly-visible PDB-101 banner signalled an enhanced commitment to developing educational resources. All of these resources were integrated within the main RCSB PDB website architecture with the intent of fostering even tighter connections and interactions between the educational content and the scientific data principally consumed by researchers.

The PDB-101 logo was inspired by a classroom chalkboard. The logo appeared on all RCSB.org website pages.
PDB-101 for the Next Generation
Creation of a better home for RCSB PDB’s educational materials worked, almost too well. Soon, the design of the initial launch wasn’t flexible enough to handle all of the new content being created: curricular materials, videos, paper models, posters, and more. The architecture of the website was limited to browsing; searching was not an option. To better-engineer PDB-101, the concept of an “E-Z PDB” was revisited in 2015.
In the redesigned system (that is still in use), all materials can be searched by assigned keywords or browsed using an expanded series of categories (ndr: see here) . The PDB-101 home page is configured to provide quick access to popular features. It is also designed to highlight new resources and outreach projects, including an annual Video Challenge for High School students built around PDB-101 materials and a biomedical theme. SciArt galleries feature work by David Goodsell and molecular illustrator pioneer Irving Geis (1908-1997).
PDB-101 today is an evolving tool for telling stories about structural biology and the PDB archive. While rigorous analysis of the impact of these resources still remains a challenge, feedback from in-person activities, reprint requests, and website analytics confirm that PDB-101 content is used in classrooms around the world.
Rapidly providing educational materials that help explain the structural biology behind public crises has helped build up our audience. In 2020, PDB-101 hosted >850,000 users and hosted >2.6 million page views. Much of this traffic was directed towards coronavirus-related content during the first half of the year. We experienced similar surges in PDB-101 traffic when we provided materials in a timely response to Ebola in 2014 and measles in 2019.
Molecular animations and videos are also very popular, with most of the viewer traffic served by third party resources. The updated What is a Protein? video, for example, has been viewed >1.6 million times on YouTube.
Continued Celebrations
We at RCSB PDB are enjoying the golden anniversary celebrations by adding unique content to PDB-101, including Structural Biology Playing Cards (here), a historical video (here) that highlights PDB-related experimental techniques, and a PDB50 board game (here) .
Acknowledgements
PDB-101 would not exist without RCSB PDB members past and present, especially: Cole Christie, Shuchismita Dutta, Sutapa Ghosh, David S. Goodsell, Rachel Kramer Green, Brian Hudson, Cathy Lawson, Yuhe Liang, Robert Lowe, Ezra Peisach, Irina Persikova, Chris Randle, Monica Sekharan, Chenghua Shao, Yi-Ping Tao, Maria Voigt, Jessie Woo, Jasmine Young, and Christine Zardecki; and the guidance of RCSB PDB Director Stephen K. Burley and Director Emerita Helen M. Berman.
RCSB PDB is funded by the National Science Foundation (DBI-1832184), the US Department of Energy (DE-SC0019749), and the National Cancer Institute, National Institute of Allergy and Infectious Diseases, and National Institute of General Medical Sciences of the National Institutes of Health under grant R01GM133198.
PDB-101 Highlights
- Coronavirus Resources
- Guide to Understanding PDB Data: Small molecule ligands
- Molecular Coloring Books
- Irving Geis Digital Archive
- Award-winning PDB-101 Resources
Related Resources
