CSD-CrossMiner in Action: Identifying a Target Protein for New Pharmaceuticals via Scaffold Search
Here we highlight a recent paper in which CSD-CrossMiner was used to find potential target protein(s) for newly synthesized spirobarbituric scaffolds. Such compounds have promising biomedical applications as anticonvulsants, anti-AIDS agents and anti-inflammatory remedies.
This is part of our series highlighting examples of the Cambridge Crystallographic Data Centre (CCDC) tools in action by scientists around the world.
Summary
In this work, researchers at the N. D. Zelinsky Institute of Organic Chemistry designed novel one-pot multicomponent reactions to synthesize unsymmetric spirobarbituric dihydrofurans – a promising new scaffold with many potential pharmaceutical applications, including in the treatment of various inflammatory, infectious and immunological diseases.
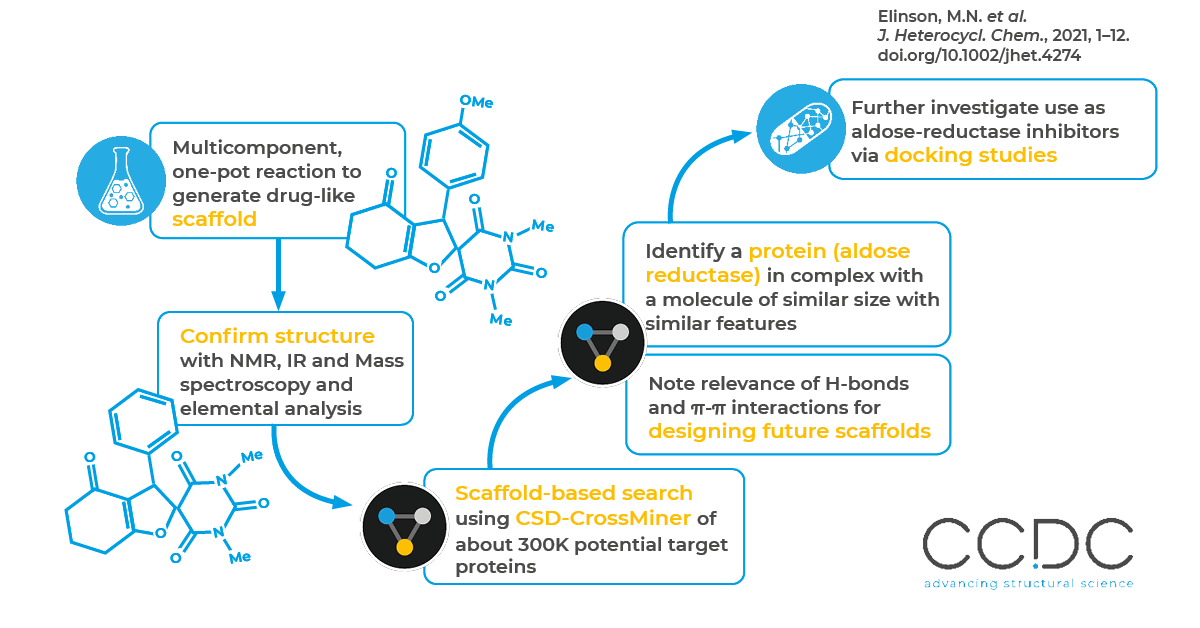
Once the spirobarbituric dihydrofurans were synthesized and the 3D structure obtained, the researchers looked for potential protein target(s) by performing a scaffold-based search using CSD-CrossMiner. They searched nearly 300,000 structures in the CSD-CrossMiner Protein Database (PDB) subset. Among the structures acquired from the scaffold search, they selected human aldose reductase in complex with a small molecule. Along with identifying a target protein, CSD-CrossMiner revealed the similarity of the co-crystallized ligand and the spirobarbituric dihydrofuran compound – highlighting the role of hydrogen bonds and π-π interactions in the protein-ligand. Molecular docking studies supported the scaffold search findings, further suggesting the promising application of spirobarbituric dihydrofurans as an aldose-reductase inhibitor over aldose reductase–mediated diseases and diabetes in particular.
Why
Privileged structures, or scaffolds, have become a common goal in the search for pharmaceutically active compounds. Scaffolds are compounds that likely bind to the same target protein but contain different core structures that present new pharmaceutical applications. Most scaffolds have a rigid heterocyclic system with a special orientation of functional substituents for target recognition. Spirobarbiturates are sought after scaffolds because of their broad spectrum of biological properties. They can act as a tumour necrosis factor-alpha (TNF-α), and they are used in the treatment of various inflammatory, infectious, immunological or malignant diseases. Spirocyclic (pyrimidine and thiadiazole) derivatives also exhibit potent antimicrobial activity.
Once a new scaffold is synthesized, it must be paired with a target protein for pharmaceutical use. However, the search for target proteins for newly synthesized compounds is non-trivial. As the researchers point out: in 2020, there were 163,633 registered, preorganized structures in the Research Collaboratory for Structural Bioinformatics Protein Data Bank (RCSB PDB). This makes a non-automated, manual analysis and enumerative search of a protein-ligand pair unreasonable. Instead, the researchers used CSD-CrossMiner to find an effective co-crystalization with aldose reductase.
How
First, the researchers optimized a one-pot, electrocatalytic multicomponent approach to synthesize substituted unsymmetric spirobarbituric dihydrofurans in 62%–76% yields. They used CSD-CrossMiner to complete a scaffold search for a target protein that could dock with the newly synthesized compounds. They identified aldose reductase with a co-crystalized ligand where the position, orientation and amount of hydrogen bonds and π-π interactions of the co-crystal ligand and the spirobarbituric dihydrofurans were very similar. They then confirmed the findings via molecular docking experiments.
Read more
Read the full paper: “Electrocatalytic multicomponent one-pot approach to tetrahydro-20 H,4H-spiro[benzofuran-2,50-pyrimidine] scaffold,” J. Heterocycl. Chem., 2021, 1484–1495.
Learn more about CSD-CrossMiner.
Read how CSD-CrossMiner fits into a computer-aided drug discovery workflow via CSD-Discovery.
Explore more examples of CCDC tools in action.
Download the Infographic CrossMiner: Drug Discovery through Data Mining