Top 200 Drugs in the CSD
In recent years, the Njarðarson research group at the University of Arizona have created posters of the top 200 drugs by sales for education, research purposes and scientific communication.1 When the latest poster based on 2018 data became available for download on their website, I was interested in finding out how many of the top-selling drugs have crystal structures in the Cambridge Structural Database (CSD).2
Looking more closely at the top 200 revealed that 76 drugs are large biological entities, more suitable for inclusion in the Protein Data Bank (PDB), rather than the CSD.3 Interestingly, this included 7 of the top 10 drugs, and shows an increase from 50 of the top 200 pharmaceuticals in 2012. The remaining 124 small molecule drugs were found to comprise 131 unique compounds, and the database could be searched for structures of these compounds using the CSD Python API. To make use of the API’s similarity search functionality, I downloaded SDF files for each compound from either DrugBank or PubChem, and used these as the input for a python script to search the CSD for similar compounds with a Tanimoto similarity score of at least 0.7 (a score of 1 indicating an identical compound).4,5
Next, I set about checking the structures found by the script, and, where multiple structures of a compound were present in the database, selecting the most suitable structure for inclusion on our own version of the poster. For compounds without any hits, I double-checked the CSD using WebCSD to search for compound names, and also checked online for any data that may be in the public domain that we could then add to the database, which resulted in several additional structures. Overall, of the 131 unique drug compounds, 91 were found in the CSD, meaning that 93 of the 124 small molecule drugs have a crystal structure of at least one of their component compounds. Given the current impact of coronavirus around the world, it is worth noting that two of these drug compounds, darunavir and sofosbuvir, have been identified as potential candidates to target COVID-19, and have crystal structures in the CSD.
With the crystal structures available in the CSD identified, I began work on creating our own version of the poster, with colour coding to show which drugs were small molecules with crystal structures in the CSD (green), which were biological agents falling outside the CSD inclusion criteria (grey), and which were small molecules whose crystal structure has not yet been determined or deposited in the CSD (white). Mercury, CCDC’s visualisation software, features the ability to create POV-Ray raytraced images, and this was used to generate high-quality 3D images of the drug crystal structures in the database.
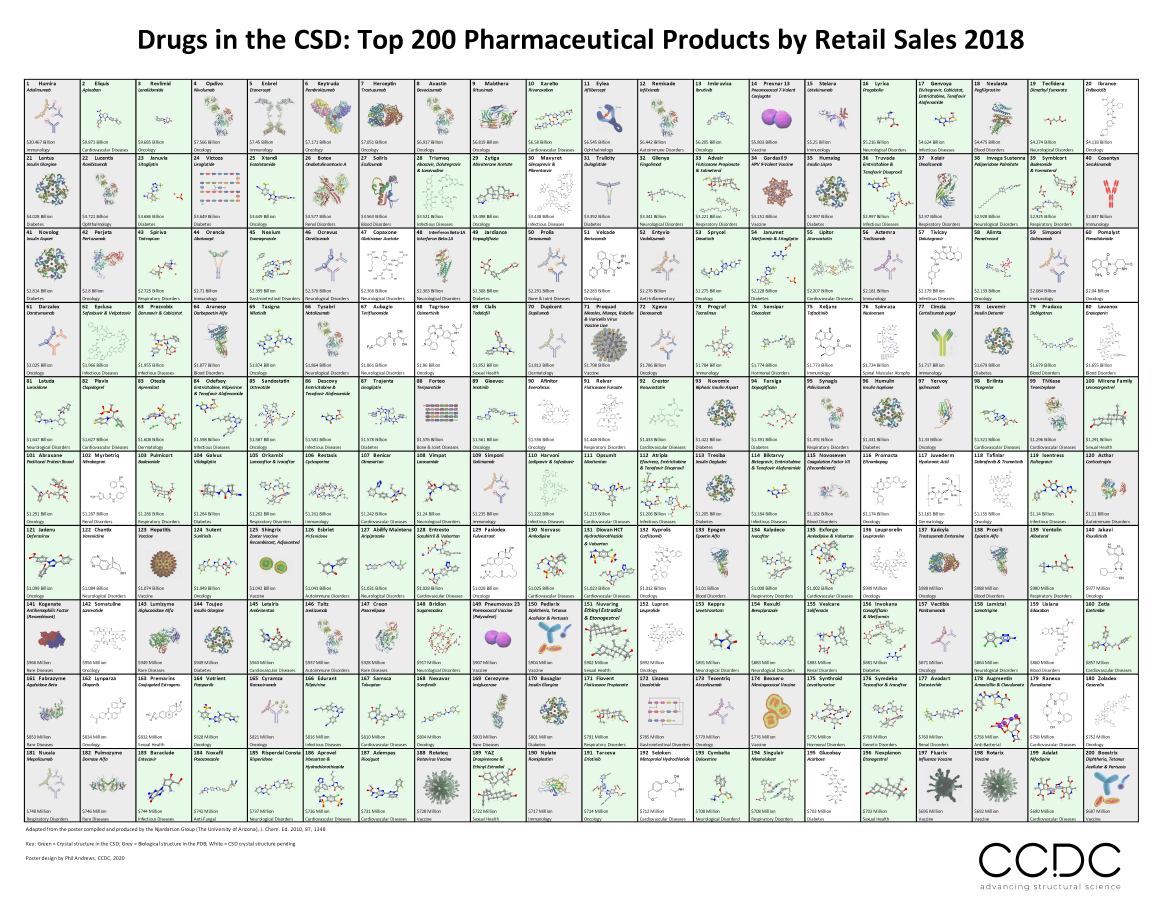
Image to show the new CSD Drugs Poster which you can download at the bottom of this blog
As you can see from the poster above, there are still a few compounds we don’t have in the CSD – this is obviously a situation we would like to fix! It can sometimes be challenging to obtain data for pharmaceutical compounds due to patent protection of intellectual property, and where diffraction data is included in patents, often only cell values or a powder pattern are available. Nevertheless, we are busy searching patents and publications and will be contacting the relevant pharmaceutical companies if appropriate too. We hope that data for these missing compounds will be deposited with us soon, and we can keep the poster up to date with even more of the top 200 drugs in the CSD.
A high-resolution PDF file of the poster is available to download here.
- A Graphical Journey of Innovative Organic Architectures That Have Improved Our Lives, Nicholas A. McGrath, Matthew Brichacek and Jon T. Njardarson, Chem. Educ., (2010), 87, 1348-1349, DOI: 10.1021/ed1003806
- https://njardarson.lab.arizona.edu/content/top-pharmaceuticals-poster
- https://www.rcsb.org/
- https://www.drugbank.ca/
- https://pubchem.ncbi.nlm.nih.gov/