Keeping up the pace of change
We announced in February a substantial change to our software release pattern with a target of four software releases during 2018, rather than the usual single release around November time. This shorter development and release cycle allows us to be more responsive to user feedback as well as helping to ensure a continuously improving and stable software system. This current update is our second software release of the year (2018 CSD Release Update 2) with two more planned in the second half of 2018.
Our work on the underlying technology of the CSD system to evolve the database and search software has been continuing since the last update. This release includes completely rewritten text searching functionality within ConQuest which has clarified and simplified the behaviour behind the scenes as well as fixing a range of long term issues. We expect to also incorporate rewritten 2D and 3D search functionality in the next release (Update 3) which will bring consistency to the searching across all our software (including Mercury, ConQuest, WebCSD and the CSD Python API).
We have also been looking at the usability of the Hydrogen Bond Propensity (HBP) functionality within Mercury for CSD-Materials users and this update sees a re-design of the display of results at the end of the process. The HBP wizard takes the user through the process of building a knowledge-based model for a target compound to predict the most likely hydrogen-bonding outcomes for that system. When viewing the results, the interface now presents all the information the user needs to assess the possible hydrogen-bonding networks (along with any observed structural data) in a single page.
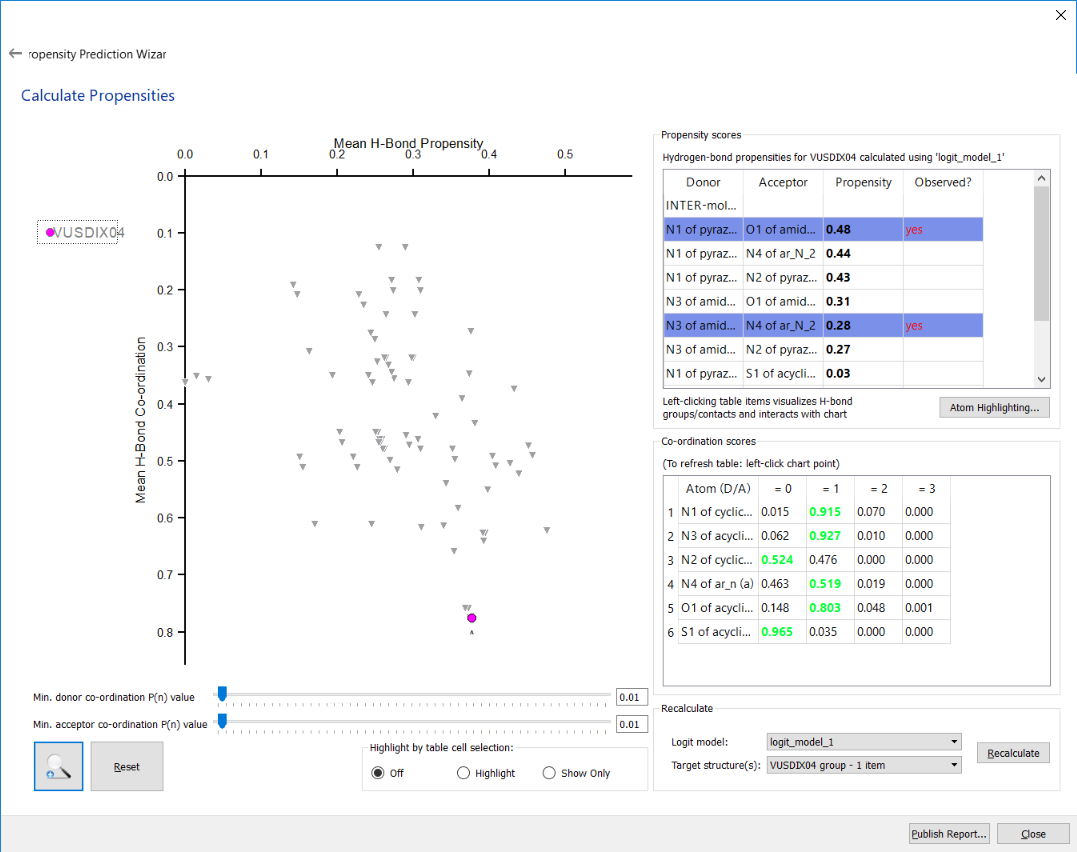
The results of a Hydrogen Bond Propensity (HBP) analysis for CSD refcode VUSDIX04 (polymorph XLI of axitinib)
As always, to find the latest software and data updates, click on Help > Check for Updates in either Mercury or Hermes. This will auto-update your system, but if this doesn’t work, you can go to the CCDC Downloads Page and download the updates manually. We are aware that users on Mac OS, or using proxies, must update manually for now and we apologise for this inconvenience. We believe that this software update will fix those issues, such that Update 3 should be straightforward for everyone to apply using the auto-update process.
Of course, we have made a range of other general improvements to the system to improve robustness and reproducibility. These include ensuring more consistency in IsoStar contour plots and providing an easy link in the Mercury (& Hermes) CSD Python API interface(s) to the output directory. CSD-CrossMiner has been updated to include better reference molecule loading, and to allow numeric range filtering of results. In addition, CSD-CrossMiner is now auto-updateable for future releases. We’ve also worked hard to improve the user guides for several of our CSD-Discovery products.
Alongside these improvements to the software, the CSD itself continues to expand. With the latest data update, the number of entries in the CSD has now passed the milestone of 950,000. We continue to be on course to reach a million structures during 2019! To find out more about the latest data update, see the 2018 CSD Data Updates web page.
As always, please do get in touch if you have any questions or comments about the CSD data or software – you can reach us via e-mail at or through our CCDC social media channels ( Facebook and Twitter).
Pete Wood, Product Manager for the CSD System
Tags