2018 CSD Release: Ellipsoids and data mining
It’s beginning to look a lot like Christmas and that means it must be time for the annual CSD Release! The 2018 CSD Release is indeed now available for download from the CCDC website – do fetch the new installers if you haven’t done so already. This year’s release includes a range of improvements in the software as well as the data based on your feedback over the year, including one improvement that has been very frequently asked about in recent years.
Thanks to a lot of hard work on the underlying technology of the system, which will continue during 2018, this year marks the first time that anisotropic displacement parameters (ADPs) have been released as part of the CSD! These ADPs (sometimes referred to as thermal ellipsoids) capture the average deviations of atoms away from their refined positions in the crystal structure model and can give valuable insight into aspects of the structure such as thermal motion and static disorder. We’ve had requests from users about these for many years now and this release makes the additional information available for display in Mercury for over 220,000 entries. The data can also be accessed using the CSD Python API, as well as through a new subset in ConQuest – the ADPs available subset. We hope that this will open up a wide range of options for more detailed analysis of structures making use of the ADP information.
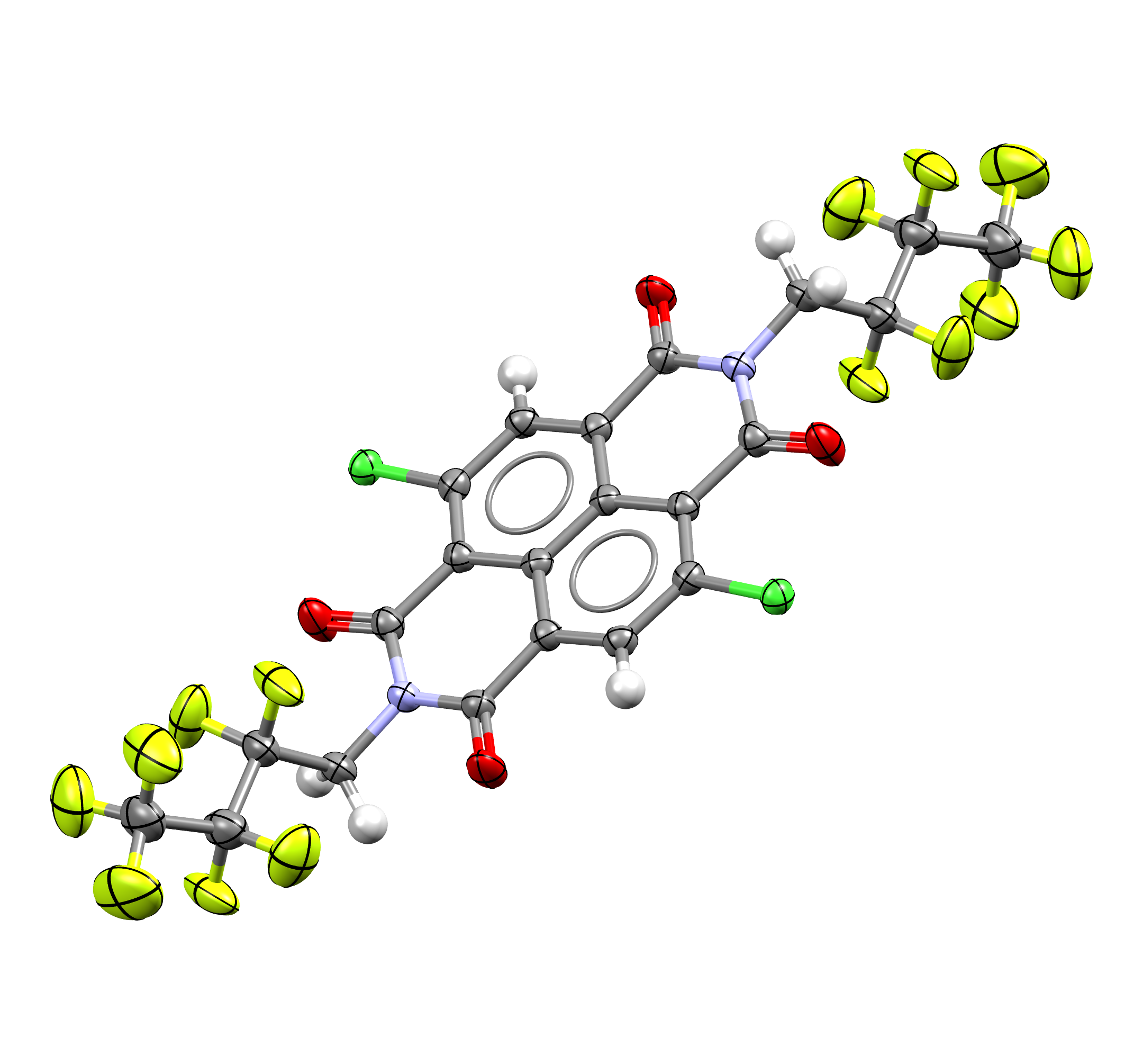
An example of a CSD entry with ADPs now available, a dichloro-naphthalene diimide structure (refcode: BUYJAH01, DOI: 10.5517/cc135btf)
We’re also introducing a new application this year for interactive pharmacophore searching; CSD-CrossMiner. This new addition to the CSD-Discovery suite provides a flexible and responsive environment for searching against both small-molecule crystal structures and protein-ligand binding sites. The CSD and the PDB both provide sources of information for new ideas; for example, what functional groups bind to similar pockets? What types of chemistry can act as similar molecular scaffolds? We can explore these questions using CSD-CrossMiner which allows the user to search for similar patterns based on pharmacophoric or sub-structure features as defined by the user. It’s always been possible to investigate patterns in the CSD using ConQuest, but now with CSD-CrossMiner, the user can interact with the search in a much more intuitive way, reducing the barrier to use.
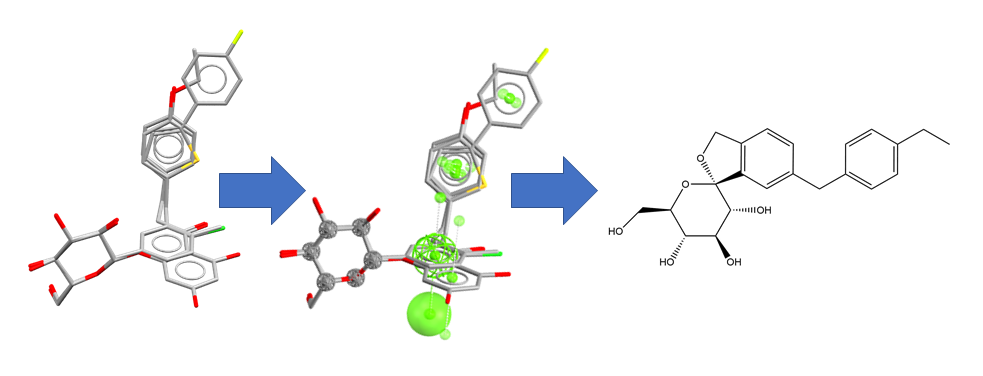
CSD-CrossMiner: Structure data to pharmacophore query to hits interactively (see Oktake et al. 2012 for the science that inspired this example)
Other improvements in the 2018 CSD Release include updates to the database itself – over 60,000 new entries have been added this year, bringing the total size of the CSD to over 900,000 entries for the first time. This includes a record breaking 5,247 new CSD Communications – structures solely published through the CSD. The database has also been enhanced with targeted improvements to over 62,000 entries, including the addition of oxidation state information to over 21,000 entries. CSD-System software enhancements include new bromine & iodine probes added to Full Interaction Maps included in the CSD-Materials suite, whilst CSD-Discovery & CSD-Materials users will benefit from an improved CSD Conformer Generator which more effectively handles sparse distributions and rare ring conformations. For further information about the 2018 CSD Release, see What’s new for 2018 as well as the respective user guides.
We hope that you enjoy the new features in the release this year. Do get in touch to let us know any feedback that you have about the 2018 CSD Release through , or our CCDC social media channels (Facebook and Twitter). Any comments or questions from our user community are always welcome!
Pete Wood, Product Manager for the CSD System