Not so Weird and Wonderful?
One of the benefits of my role at the CCDC is the chance to look at some of the latest scientific research taking place, as I review structures before they are added to the Cambridge Structural Database (CSD). Occasionally I come across a structure that looks quite unusual at first glance, so much so that it’s hard to resist taking a closer look.
Recently this occurred when I saw OGUROZ (http://dx.doi.org/10.5517/cc1j0djg). I’ve attempted to create a clear representation of the structure in the image below; if you think it looks complex that’s due to the 56 independent molecules in the asymmetric unit of the crystal. This number of molecules per unit cell (known as Z’) is the largest reported to date and is described in a recent Chemical Communications paper http://dx.doi.org/10.1039/C5CC04219D.
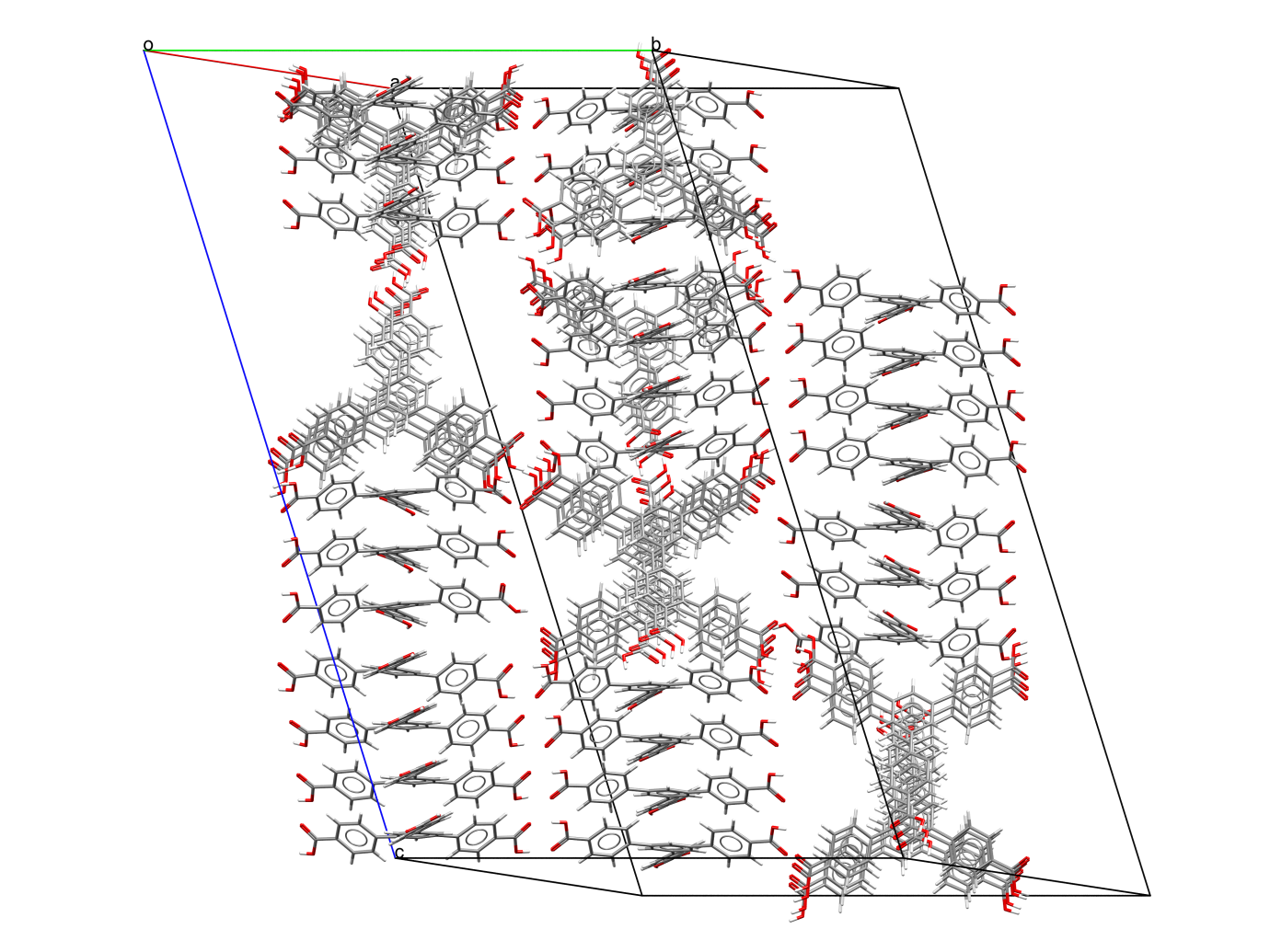
Unit cell of structure OGUROZ, showing 56 independent molecules
Every year, tens of thousands of entries are added to the CSD, each containing details of the publication and the chemistry and crystallography of the structure. Our automated systems and the watchful care of editors combine to ensure the weird and wonderful structures we see really are as weird and wonderful as they first appear, but maintaining a consistent representation over the structures in the CSD is always a challenge. This year we’ve been celebrating the CSD’s 50th anniversary, so our goal is not only to represent the CSD entries we add today in a consistent way, but also to ensure that the legacy of entries we’ve maintained over the last 50 years are equally consistent.
Each year we issue a new version of CSD-System. This contains the entire CSD, including all structures which have been deposited since the previous year’s release. We take this opportunity to review the complete CSD, and that process has already started for this upcoming year’s release. However, this time around is significantly different as we have the additional benefit of the recently released CSD Python API (link to http://www.ccdc.cam.ac.uk/News/List/post-37/) to help our work.
So, to go back to the structure OGUROZ, it’s well known that high Z’ values in crystal structures are unusual, and a group in Durham has looked at this in detail (link to http://zprime.co.uk/database). By writing a short Python script, we can use our API to easily find all such high Z’ values (see the graph below), and target our checks and scientific expertise on CSD entries where we believe they’re needed most. You can help too, of course, if you happen to find any unusual CSD entries you’d like us to investigate, you can tell us by emailing
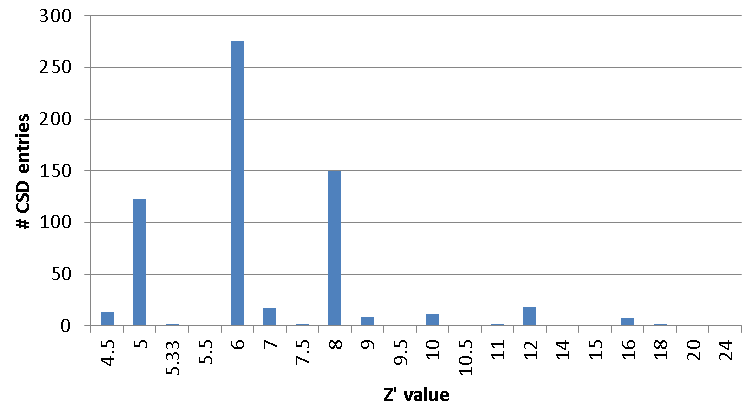
Graph showing a subset of CSD entries listed with Z’ > 4
The graph below shows another example of the kinds of checks we can now carry out. The graph shows the measurement of void space in a subset of the CSD containing around 100,000 organic crystal structures. Using the existing search capabilities of ConQuest and the new features of the API makes it simple to choose specific subsets of structures for investigation. Void space calculations are not part of the data we receive from depositors, but by using the API we can harness the tools in Mercury to run these checks for us over this large dataset.
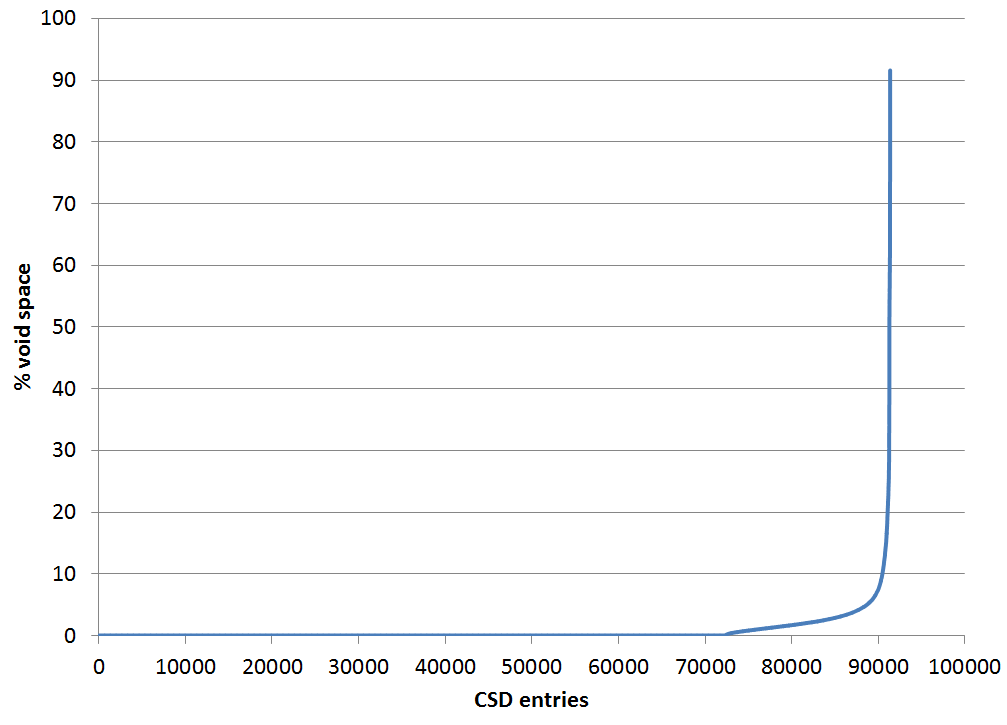
Results of the void space analysis of around 100,000 organic CSD entries
You can see that the vast majority of this subset of organic structures contain, as you would expect, no solvent-accessible void space. Around 20% of structures contain a small degree of void space, perhaps indicative of non-modelled hydrogen atoms or some disorder in the structure. The graph does have a rather dramatic tail, and this refers to around 200 of the almost 100,000 structures that really do contain significant voids. We can review each of these CSD entries in turn and ensure the comments and remarks we add to CSD entries help make users aware of the issues with these structures.
With the new tools we have available it’s much quicker and easier for us to review all aspects of the entries in the CSD. Of course we’re keen to respond to suggestions from our users too; we’d be interested to know how you would like to see the CSD enhanced and improved in the future.
These latest API-driven improvements are good news for us, and good news for you too, as it means when you search the CSD you can be confident in the fact that you are receiveing accurate and consistent results from all 50 years of crystal structures. I think that’s a pretty good birthday present for everyone!
Seth Wiggin
Senior Scientific Editor
Tags
CSD (153)
CSD System (49)